|
Redbiom can be used to identify public data within Qiita for a meta-analysis focused on some particular factor (or factors). Issues that only occur with Vulkan should be logged in the Dota-2-Vulkan tracker.This documentation provides an introduction to accessing and processing public data within the Qiita database for re-analysis. If you have a general issue or non-system-specific feature request please go to dev.dota2.com. Tracker for issues specific to Linux and Mac in the Reborn client.
Qiime 2 Help Requested For Activate How To Search ForAs this requires only a relatively small data set, and the question is of a less exploratory nature (than the second example), it will be possible to carry out the entire process for this first example within Qiita.The second example poses a meta-analysis type question about the microbiome which has not yet been fully addressed in the literature: does a person’s frequency of exercise affect their microbiome? To answer this type of exploratory meta-analysis type question does not necessarily require a new study, it may be possible to re-purpose publicly available data for the analysis. This data can then be used for informing and justifying clinical trial format or experimental set-up. Both datasets are used in the first section ( Retrieving Public Data for Own Analysis) while Processing Public Data Retrieved with redbiom uses the AGP data and the Statistical Analysis to Justify Clinical Trial Sample Size Tutorial uses the data used in Casals-Pascual et al 2020.This tutorial will use two different datasets to highlight both the different questions that can be asked with existing, open source data, and the different methods that can be used to do so.The analysis of clinical microbiome data, selected for its similarity to the type of samples one plans to collect for one’s own study, allows one to produce example data before starting such a study. QIIME runs under Linux or Mac OS X, but not under Windows.To illustrate this utility, the following tutorials provide guidance on how to search for and download data using redbiom ( Retrieving Public Data for Own Analysis) and how to process data retrieved with redbiom using QIIME 2 to allow (re-)analysis of said data ( Processing Public Data Retrieved with redbiom).Furthermore, the third section (the Statistical Analysis to Justify Clinical Trial Sample Size Tutorial) provides guidance on a specific example of how public data may be repurposed, in this case to justify clinical trial or experimental sample size.Two datasets are used in this documentation:The data used in Casals-Pascual et al 2020 (originally from Halfvarson et al 2017).See Introduction to the example data for more detail. Introduction to taxonomic analysis of amplicon and shotgun data using. This may include data from studies with completely different research goals, which one would otherwise have been unlikely to realize could be used.2.A search for IBD brings up 7 studies, including Study 1629. In this case, the Redbiom plugin can be used to search by metadata attributes e.g. However, in most scenarios, one would not know of a specific study to search for beforehand. On the Qiita website select the Study -> View Study option, specify metadata in the tab down menu next to the search box and search for 1629 to find the data for this first example. This will be demonstrated by finding the data used in Casals-Pascual et al 2020, which cites that the data used was retrieved from a study in Qiita with ID 1629. This is a larger study and will require the use of the command line version of redbiom, and coding tools beyond Qiita.The following section will explain how to retrieve the data for these examples, beginning with the simpler clinical data example.Retrieving data using the redbiom plug-in within Qiita ¶It is possible to search for data and studies directly on Qiita using the redbiom plug-in.
The sample information tab includes a list of all the metadata features (e.g. Selecting data types, and then the type of interest shows a diagram of the processing pipeline, further information, and list of samples within the dataset of that type. All QIIME 2 maps and BIOMs, as well as EBI accession numbers and sample information can be downloaded directly from this main page. These selected artifacts can then be used to create an analysis as will be outlined subsequently.When browsing public studies to find appropriate data, more information on any study can be accessed by selecting it this will open a new page which includes the study abstract and details as well as options to download the study data. All artifacts of one type can then be selected by using add all, or specific artifacts can be selected using per artifact -> add. Selecting the green plus for the study of interest reveals the study data, in the form of Qiita artifacts. This opens a window where one can view all selected artifacts and exclude blocks of samples or individual samples that are not required/relevant (first select a particular artifact to view these). Once the artifacts have been added select analysis from the top bar and, from the drop down menu, select create from selected samples. Perusing these features may give a better indication of whether a study can be repurposed for one’s own analysis.A search using the redbiom plugin within Qiita therefore allows one to either download data for study on another platform or to select artifacts for processing and analysis within Qiita. Hp recovery flash disk utility for windows xpThis is also possible for another’s analysis as long as it is public. After processing raw data with Qiita the artifacts can be downloaded by selecting an artifact and clicking ‘ generate a download link’. The Statistical Analysis to Justify Clinical Trial Sample Size Tutorial) explain how to process the raw data that you have just selected within Qiita for analysis. Note that you should should select “ Merge samples with the same name - useful when merging multiple preparation artifacts” if you want to use the metadata from the studies the samples originated from.Other tutorials (e.g. While artifacts, or raw study data, found within Qiita can then be downloaded after accessing the study and finding their ID, the command line version of redbiom allows searching and direct download all within the command line. The following notation can be used to fetch:Where / should be replaced with the appropriate study-id/prep-id.Retrieving data with redbiom in the command line ¶While the redbiom plugin for Qiita is useful for simple searches, and when finding data for processing and analysis within Qiita, the command line version of redbiom has increased functionality, and is particularly useful when data will be processed outside of Qiita. To fetch specific types of data one can also use specific calls. This is also useful in the situation where one wants to use the processed artifacts from a public study (simply click on an artifact in any public study’s processing network to view its artifact ID). 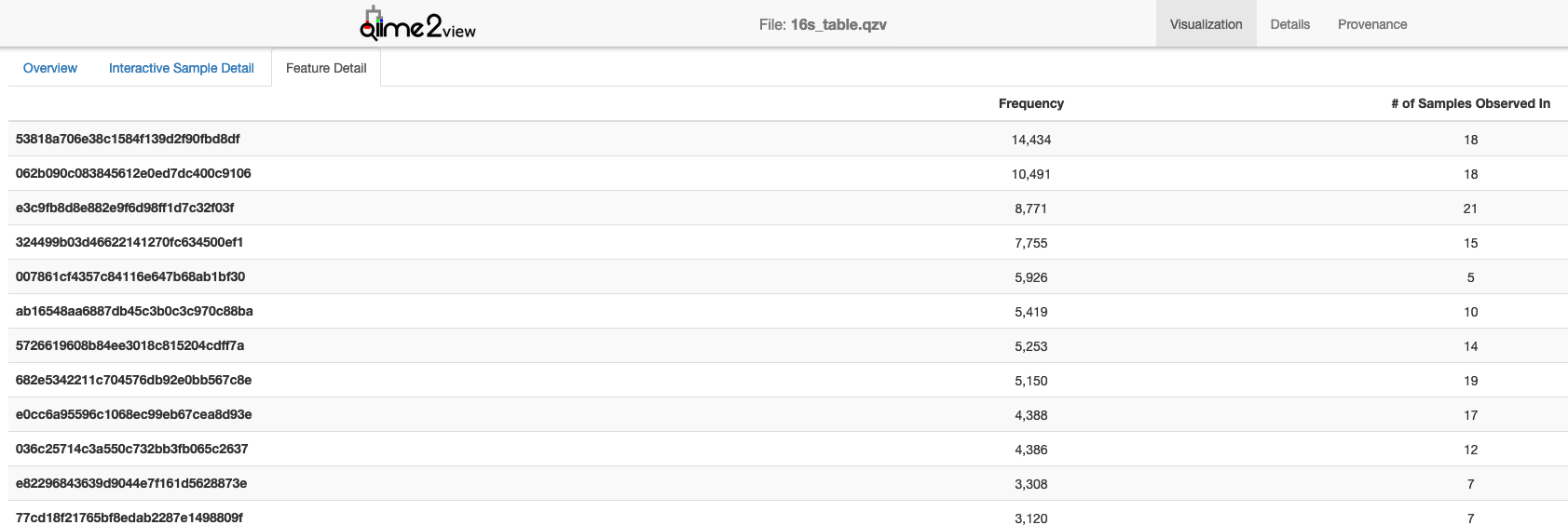
It accepts both ‘natural language’ (but uses stemming) and python-like grammar (separately or in combination). Metadata is particularly useful for our purposes. Taxon will return features associated with a taxon. Both of these commands require a specified file and so are not as relevant to initial exploratory searches. , =>, = to see the categories which include that keyword, one can then learn more about a specific category with summarize metadata-category -category -counter.Querying and fetching sample data using Redbiom requires a specified context ( -context ).
0 Comments
Leave a Reply. |
AuthorJoseph ArchivesCategories |
 RSS Feed
RSS Feed
